I came across brilliant review by Jens Walter et al. that resonates with my thoughts in microbiome field. I also work in the same field doing similar experiments, but sometimes it is better to stop and look back what you have been doing. This review has discussed some important points on the transmissibility of human diseases in human microbiota-associated mouse models. They acknowledge the importance of human microbiota-associated mouse models as valuable resource in the microbiome field. However, through systematic review they demonstrate that according to current literature it is easily possible to transfer (any!) human disease in mice through microbiota transplant. The inflated positivity and hype around these experiments must be criticized. Most importantly, they talk about the relevance of negative transplant studies, which are seldom published, and urge to use rigorous statistics to improve the data interpretation. Following are some excerpts from the review, though reading full review is highly recommended.
The problem with the concept:
“Because a “normal” or “healthy” microbiome has not been defined yet, we use the term dysbiosis to refer solely to an altered microbiome associated with a specific disease or condition as compared to a control. Whether such “dysbiotic” microbiomes are the cause or consequence of disease, or whether both are caused by a third factor, is, in most instances, unproven.”
“There are two well-documented diseases in the microbiome field that link a microbial biomarker with causation in disease: Helicobacter pylori-associated peptic ulceration and gastric cancer, and Clostridium (or Clostridioides) difficile infection-associated diarrhea. However, causal inferences between complex microbiomes and other inflammatory, metabolic, neoplastic, and neuro-behavioral disorders have been neither compelling nor conclusive.”
“A substantial proportion of the taxa that comprise the human gut microbiome fail to colonize in the recipient animals. Most importantly, the ecological factors (such as diet, lifestyle, disease phenotype, and human genotype) that have driven the dysbiosis in humans in the first place are absent in the recipient rodents, making it unlikely that disease-associated alterations are replicated. Taxa that colonize at sufficient levels may not engage in the EVOLUTIONARY routed host-microbe interactions that they establish with their native host.”
“Consider H. pylori and C. difficile, two members of the human microbiome with established causality in human disease, neither of which exhibit sufficient colonization to cause reliable pathology in conventional murine models.”
Results from systematic review:
“Of 38 studies meeting the inclusion criteria, all but two (36/38; 95%) concluded that fecal transfer from diseased donors resulted in at least one phenotype of the human disease being greater in the recipient mice when compared with transfers from healthy donors.”
“Of all the studies, 63% (24/38) of the studies tested for a dysbiosis in the original human donor samples, and only 29% (11/38) confirmed that at least some aspect (reduced diversity, ecosystem processes/services, loss or bloom of specific taxa, or shifts in metabolic capacity) of the “human dysbiosis” was replicated in the recipient animals. Consequently, the majority of studies did not attempt to identify the “causal component” of the microbiome. Only a few studies (34%; 13/38) identified potential underlying mechanisms linking the dysbiotic microbiome with disease.”
Issues with statistics
“Often, a small number (1–5) of human donors is used per human disease and control group, and these “donor microbiomes” are then replicated in a larger number of individual mice, which are then often used for statistical inference.”
“The result is “pseudoreplication,” which artificially inflates the sample size, the chance for “false positive” findings, and the evidence for a scientific claim. This problem is amplified if donor samples are pooled before the inoculation of groups of rodents, which is common practice for both the “diseased” and healthy control samples. We found that 84% (32/38) of the published studies on HMA murine models used the individual animals as the statistical inference, although animal numbers were much higher than the number of human donors and therefore inflated. Only 16% of studies (6/38) were clear in their use of the individual human donor samples as the “N” for statistical inferences or used the same number of rodents as donors.”
“When virtually all reports are positive, as in our systematic review, the greater the doubt will be and the more justified the scrutiny should be.”
Suggestions for alternative experimental approach
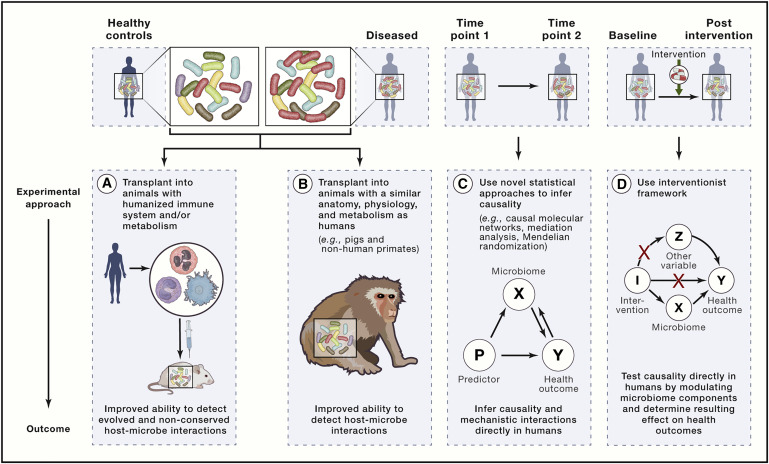
Why is it important to change?
“Specifically, the correction of altered microbiomes, be it by FMT, live biotherapeutics, or microbial products, will be beneficial only if the alterations are causal or contributory to the disease rather than a bystander response. Causal claims in the microbiome field are often overstated and disproportional to the experimental evidence from which they are derived, and it has become difficult to identify a disease state in which the microbiome has not been implicated in the scientific literature, a notion amplified in the popular press.”
“Sadly, most researchers and reviewers in the microbiome field, and almost all editors of high-impact journals, seem to regard acceptance of the null hypothesis as a failure. As with all scientific endeavor, our field would be served better if editorial policies of higher tier journals and guidelines of funding agencies discouraged any discrimination against negative data, replications, validations of experimental designs, and statistical analyses.”

