My second (and last) day of reporting from the 4th Annual North American Microbiome Congress in Washington DC, organized by Kisaco Research. Here is a roll-out of my tweet storm from today.
Good morning! In 15 minutes I will start live tweeting Day 2 from the 4th Annual Microbiome Congress, in Washington DC, organized by @KisacoRes #MBCongress2019
I am all ready to go in the front row at the plenary session, with Larry Weiss, Amanda Kay, and Ken Blount.

Morning welcome by Larry Weiss from Persona Biome
Larry Weiss @LWeissMD, CEO and founder of Persona Biome opens this morning’s session. In this field, we have undergone a transformation. We start thinking more like a microbial community, how everything is connected.
Larry Weiss: As scientists we gather data, and build models. But models are not truth. There is so much we don’t know that we don’t know. This field is still very early.
Larry Weiss: We are intimately connected to our microbiome, but have cut connections with our environment. We are a complex process of processes.
Larry Weiss: Finally, I want to talk about “shit”. We need to treat feces with much more respect. It is not only a waste product, but the product of a bioreactor.
Larry Weiss: This science is going to transform everything. We need to develop partnerships and work together.
Amanda Kay from Synlogic and Bradford McRay from AbbVie
The opening presentation of today is by Amanda Kay, VP at Synlogic @synlogic_tx, and Bradford McRae, AbbVie Discovery Immunology @abbvie about “Collaborating with AbbVie to develop a novel class of living medicines”
Bradford McRae starts off: There are several key questions for microbiome-based drug development. Most importantly: Are changes in the microbiome the cause of disease or a secondary consequence of the disease state?
Bradford McRae: Will altering the composition/function of the microbiome change the natural history of disease? Can the composition/function of the microbiome be used to stratify patient populations and/or define future disease course?
Bradford McRae: There are many factors that influence microbiome composition and function, such as diet, exercise, stress, medications, metabolism, immunity.
Bradford McRae: Some challenges for understanding the therapeutic potential of the microbiome: Intra-individual disease and microbiome heterogeneity make it hard to understand patterns of disease.
Bradford McRae: It is also still difficult to link sequence data to function, and to understand what happens in the different compartments of the gut by just looking at what comes out at the far end.
Bradford McRae: Excellent work on altering the function of the microbiome is being done in metabolic disease, such as FMT from lean donors to patients with metabolic syndrome, which improved glucose sensitivity.
Bradford McRae: Evidence that diet may alter the course of IBD through modifying microbiome function from Schwerd et al.
(link: https://europepmc.org/abstract/med/26987574)
Bradford McRae: Future developments:
1. Moving beyond descriptive analysis to molecular pathways
2. Future deliverables for patients: diagnostics, mechanisms
3. Potential for engineered bacteria to deliver specific therapies
Amanda Kay: At @synlogic_tx, we are designing for life. We do rational design for bacterial based therapy, and collaborate with partners. We do synthetic biotic(TM).
Amanda Kay: Synthetics: genetic circuits, degradation of disease causing metabolites, production of therapeutic molecules
Biotics: bacterial chassis, non-pathogenic, amenable to genetic manipulation
Amanda Kay: In the context of IBD, we @synlogic_tx started a collaboration with AbbVie @abbvie We both brought in our key expertise, in drug development and translation into drug candidates.
Amanda Kay: Goal of our collaboration was to leverage known homeostatic functions to resolve disease and maintain health in patients
Ken Blount from Rebiotix
The next speaker is Ken Blount, CSO, Rebiotix @Rebiotix, with: “Discovering the potential of Microbiota Restoration Therapy (MRT) drug platforms for the treatment of intestinal diseases”
Ken Blount: Fecal transplant has a long history of working well, in particular in Clostridium difficile infections, but how do you perform it in a controlled and reproducible way?
Ken Blount: In a healthy gut, non-spore forming class Bacteroidia constitutes ~30% of bacteria. In our platform, the Microbiome Restoration Therapy (MRT), we want to restore a healthy gut microbiome in a patient.
Ken Blount: In C. diff infections (CDI), it is challenging to restore the foundation of a healthy microbiome. Our MRT is not a fecal transplant, but a standardized consortium of live spore forming and non-spore forming microbes.
Ken Blount: We designed several trials, phase 2 trials already done, phase 3 enrolling. We got 87% of treatment success in our initial phase 2 trial.
Ken Blount: In our second phase 2 trial, we learned that 2 doses worked as well as 1, but success rate was only around 60%. Placebo had 47% success rate (because of pretreatment with Abx?)
Ken Blount: After treatment, the patients’ microbiomes shifted towards that of a healthy population (HMP dataset). It restores Bacteroidia and Clostridia, decreasing Gammaproteobacteria.
Ken Blount: We developed the Microbiome Health Index (MHI, TM). High MHI means healthy, lower MHI is a deviation from healthy. This single number allows to better differentiate healthy and unhealthy.
Ken Blount: We are now in the middle of a phase 3 trial. We are working on a new stable formulation, in an oral capsule, RBX7455, which had a 90% success rate in a phase 1 trial.
Ken Blount: We are working on many other applications as well (for the enema formulation): ongoing trials in VRE, pediatric UC, UTI, hepatic encephalopathy.
We will now have two talks in the session “Regulation of biotech: Are we Prepared? “
Jim Weston from Seres Therapeutics
Jim Weston, senior VP at Seres Therapeutics @SeresTX will talk about: “How we navigated the evolving guidelines”
Jim Weston: I will tell you about the process we did to get our products to market and collaborate with the regulatory offices such as the FDA.
Jim Weston: Our strategy at @SeresTX is a focused R&D, (e.g. C. diff infection), and to have a great clinical manufacturing operation. How do you meet the needs of patients as well as regulators?
Jim Weston: Therapeutic Microbiome Products are regulated in US as both biological products (drugs) as well as live biotherapeutic products. The best strategy is to have case-by-case collaborations with the regulators.
Jim Weston: TMPs can vary from stool (FMT), communities of strains, single strains, or genetically modified strains. These all require different regulatory processes.
Jim Weston: In the US, we have one agency for approval, the @US_FDA. It is best to start work with the FDA in an early stage. In Europe there is the EMA @EMA_News and national level agencies.
Jim Weston: It would be great to have some guidance for (fecal) donor screening and safety. Trial design needs guidance too, but end points are well defined.
Jim Weston: How do we ensure good manufacturing, and measurement of results? Regular interaction between industry and regulatory agencies is key.
Larry Weiss from Persona Biome
The second talk in the session about regulatory agencies, will be by Larry Weiss @LWeissMD, CEO of Persona Biome, with “Re-writing the rules for microbiome therapeutics”
Larry Weiss: We are trying to get really solid data, but not by hacking the system or evading the rules. There are existing rules that we can follow, but solid science is important.
Larry Weiss: “The @US_FDA regulates two things: substances and words” – Peter Barton Hutt.
Larry Weiss: We are changing the states of pharmaceutical development, moving from pharmacology (chemistry) towards systems biology (microbiology). This is challenging a lot of our structures, including regulatory.
Larry Weiss: Shows the definitions of “drug”, “cosmetic”, “dietary supplement” by the FDA. There are several situations where it is difficult to distinguish between these categories.
Larry Weiss: The definition of GRAS: Generally Recognized as Safe states “no genuine dispute among qualified experts” – ha! When does that happen?? You can define your own product as GRAS (food ingredient).
Larry Weiss: There are many overlapping categories.
* Drug, cosmetic, medical food, probiotic?
* Bugs as drugs (probiotics, known commensals), drugs from bugs (purified extracts), microbiome manipulation (FMT, phages, prebiotics).
Larry Weiss: Each product that you might develop might have to follow a different path. It is not about evading rules, you have to make ethical decisions.
We will have a coffee break and then I will go to one of the three parallel sessions. Hard to choose! But I will probably go to the “Food and the Microbiome” session with Kristen Beck @theladybeck and Cindy Davis @NIH
Back from coffee, at the “Food and the Microbiome” session with Kristen Beck @theladybeck (we finally meet!) @IBMresearch , and Cindy Davis @NIH
Kristen Beck from IBM Research
Kristen Beck: I get asked a lot if IBM is involved in life sciences, and the answer is Yes! We have several collaborations in the microbial space, including in the microbiome. 20/3000 researchers at our division are involved in the microbiome.
Kristen Beck: We have partnerships with e.g. UCSD Center for Microbiome Innovation, and with Mars, to look at microbes in the food chain.
Kristen Beck: Food poisoning is very frequent, and companies invest a lot in processes to limit food safety hazards. There are known (e.g. Salmonella) and unknown (never anticipated before) hazards.
Kristen Beck: The microbiome will respond to its environment, much like a canary in a coal mine. In food samples, there can be microbes, which can be detected by sequencing.
Kristen Beck: We analyzed 312 terabytes of data, generating >270,000 data files, building up datasets from food microbiomes. We also build a bioinformatics tool that could work for a wide range of people, including those who cannot run from the command line.
Kristen Beck: Kiwi, an advanced prototype of food microbiome analysis – an easy to use interface for analyzing food microbiomes.
Kristen Beck *shows some Kiwi dashboard screenshots *. On the left, you see a good sample, with no microbial hazards detected (in blue and green) – on the right, hazards detected (in red and orange). These are easy to interpret.
Kristen Beck: For the more advanced users, we offer advanced analysis tools of these samples.
Kristen Beck: We work with deep sequencing of the metatranscriptome (mRNA), around 350 million reads per sample. Samples all retrieved from same factory and sample type.
Kristen Beck: With food, it is not always clear what the matrix (host) is: is it chicken or some other type of meat? This provides unique computational challenges to the bioinformatics analysis.
Kristen Beck: We search for a reference set of 6,000 plant and animals genomes, then use Kraken to filter out these matrix reads. This filtering should only remove the matrix, not the microbial reads.
Kristen Beck: We can also use this process to see the composition of host DNA in a sausage sample (55% sheep, 35% cattle, 7.5% pig and 1% horse!).
Kristen Beck: In screening chicken samples, we found most of them to be >99% chicken, but some had pig and cow DNA in them.
Kristen Beck: The microbiome analysis is then done on the non-matrix nucleic acids. We show the high abundance microbes to signal if something is wrong.
Kristen Beck: Unclassified reads are a missed opportunity, caused by lack of homology. Reference databases are biased towards cultured organisms. We are building better reference sets that include many more genomes to better capture the diversity of strains.
Kristen Beck: Introducing OMXWare: We @IBMResearch assembled 166,000 high quality genomes, annotated genes, proteins, and domains. It has lots of data in a structured and optimized DB2 database.
Kristen Beck: OMXWare can be used e.g. to extract novel CRISPR/Cas sequences and to find higher % of functional annotations.
Cindy Davis, National Institutes of Health
The next speaker is Cindy Davis, Office of Dietary Supplements, National Institutes of Health @NIH with “Diet, microbiome and health: the influence of diet on the intestinal microbiome”
Cindy Davis: Diet shapes the microbiome in humans: globally distinct populations, food pattern consumption vs enterotypes. High fat vs high fiber diets are associated with different microbiomes.
Cindy Davis: Prevotella is associated with diets with lots of fibers, while Bacteroides is more prevalent in Western diets with more animal fats. Diet can influence the microbiome composition, even in short term experiments.
Cindy Davis: Shows data from the Daphna Rothschild Nature paper.
Environment dominates over host genetics in shaping human gut microbiota
(link: https://www.nature.com/articles/nature25973)
Cindy Davis: Shows WHO definition of probiotics, which now form a >2 billion dollar sales market in the US. They are the 3rd most common dietary supplement. But what do they do on humans?
Cindy Davis: Stool samples do not reflect the whole microbiome in the gut. Some people’s microbiota resists colonization with probiotics.
Shows data from Zmora et al.
(link: https://www.cell.com/cell/fulltext/S0092-8674(18)31102-4)
Cindy Davis: There have been several studies using the effect of probiotics on BMI. Most of these are short duration, small sample size (underpowered), used variable strains, not preregistered, and not clinically significant.
They don’t prove anything.
Cindy Davis: So where is the science? The FDA has not approved any probiotics for preventing any health problem. There is some preliminary evidence for some effect of probiotics in certain situations.
Cindy Davis: Dietary fiber is associated with a decreased risk of colon cancer. They are fermented in the colon into short chain fatty acids (SCFA), such as butyrate.
Cindy Davis: Butyrate can affect proliferation of cancer cells (which prefer glucose) and increase apoptosis, thus plays role in cancer prevention.
Cindy Davis: Dietary fiber can protect the mucus barrier. In the absence of fiber, the gut bacteria start to eat the mucus layer.
* shows data from Desai et al. 2016
(link: https://www.ncbi.nlm.nih.gov/pubmed/27863247)
Cindy Davis: Dietary allicin (in garlic) reduces metabolism of L-carnitine into TMAO. See Wu 2015 (link: https://onlinelibrary.wiley.com/doi/full/10.1002/fsn3.199)
We need to think about food interactions!
Cindy Davis: *shows tiny graphs from many more papers by other groups*
Cindy Davis: There is a dynamic relation between microbes, food components, and microbial metabolites. Can your microbiome tell you what to eat?
Zeevi et al. : high interpersonal variability in glucose response in 800 person cohort.
(link: https://www.cell.com/cell/fulltext/S0092-8674(15)01481-6) cell.com/cell/fulltext/
Cindy Davis: When you are going to have lunch now, remember that you are also feeding your microbes. And they might want something different to eat than you. 🙂
This talk gave a good overview of the current state of microbiome research in the setting of diet, but did not present anything new. Would have been a better talk as a keynote/opening lecture, not in a specialized conference track.
Scott Jackson from the National Institute of Standards and Technology
We start after lunch with a talk by Scott Jackson from the National Institute of Standards and Technology @usnistgov, with “Standards for Microbiome and Metagenomic Measurements”
Scott Jackson: We are a non-regulatory agency, but often work together with the FDA to develop standards that can be used by industry.
Scott Jackson: The microbiome industry is rapidly growing, and standards are needed for diagnostics and methods. Many biases in metagenomic measurements, e.g. DNA extractions, primer choice, library prep, sequencing technologies, and bioinformatics analyses.
Scott Jackson: 5 years ago, it was “cowboy country” – different methods will give different microbiome community outcomes. We make references and standards to try to guide this.
Scott Jackson: The Mosaic Standards Challenge was launched in May 2018, made possible by Janssen, Biocollective, DNAnexus, and NIST. You can sign up for free. You process samples through your favorite method and upload your data.
(link: http://mosaicbiome.com/) MOSAIC: A Cloud-Based Microbiome Informatics Platform
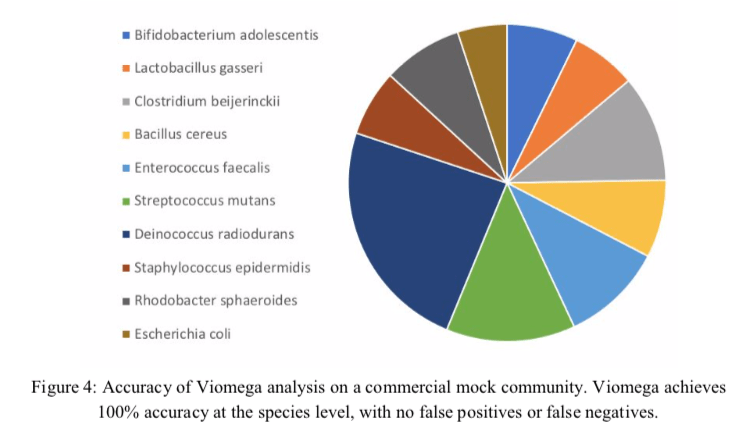
Scott Jackson: Here are some results from submitted datasets on a set of 10 bacterial standards. Results vary widely between labs, and we are looking at which factors make the most impact.
Scott Jackson: High number of reads often corresponded to more false positives (detection of strains that were not in the tube we sent).
Scott Jackson: None of the companies that do microbial diagnostics do have FDA approval, but the FDA calls on NIST for microbial standards. We have made a mix of 19 different pathogens in human DNA, which can be used to validate tests.
Scott Jackson: We made different serial dilution mixtures and tested expected vs observed abundance using 3 metagenomics analysis tools to infer specificity and sensitivity.
Scott Jackson: International microbiome and metagenomics standards alliance IMMSA provides lots of tools, workshops, videos. We also have a pathogens workshop group. Our next NIST/FDA/NIH workshop is Sept 9-10 in Gaithersburg, MD.
(link: https://microbialstandards.org/) microbialstandards.org
Moved to parallel session 1, where I caught the summary slide of Peter Karp @SRI_Intl talk, in which he presented:
* Multi organism Metabolic Route Search
* BioCyc Databases Combine
* Pathway Tools software
Curtis Huttenhower from Harvard University
The next talk will be by Curtis Huttenhower @chuttenh from Harvard University @harvard, with “Integrating molecule measurements of the microbiome for translation in population health”
Curtis Huttenhower: Our group works on methods development to analyze the microbiome on different levels, including very large-scale studies.
Curtis Huttenhower: BIOM-Mass: Biobank for Microbiome Research in Massachusetts. Developing a room temp-ship-able kit for stool and oral samples suitable for metagenomics, metatranscriptomics, metabolomics, and culture.
Curtis Huttenhower: Phase 2 of the Human Microbiome Project, HMP2, or integrative HMP (iHMP). Our lab is studying the microbiome of inflammatory bowel disease (IBD) patients over time, including CD, UC. Roughly every 2 weeks + blood draws and biopsies.
Curtis Huttenhower: This allows us to connect host response to microbiome profiles, at roughly the same timepoints. Database and raw data available here: (link: https://www.ibdmdb.org/) ibdmdb.org
Curtis Huttenhower: We can see which microbial metabolites (as measured biochemically) are enriched or depleted in IBD patients vs healthy controls and connect these to variations in microbial taxa.
Curtis Huttenhower: We can also link microbial function to strain specific phenotypes in IBD. *shows heatmap of Ruminococcus gnavus genes and their abundance in patients/controls*. Most gut microbial genes are unfortunately still of unknown function.
Curtis Huttenhower: Type 1 diabetes infant cohorts in Finland, Estonia, Russia / TEDDY study: functional shifts in genes at birth, at year 1, at year 2. Some are linked to specific functions, e.g. HMO utilization in infants in Bifidobacterium longum strains.
Curtis Huttenhower: Nicola Segata: assembled 150,000 genomes from 10,000 metagenomies, 5,000 species-level genomic bins (SGBs). Many of these genome bins are novel, many from uncharacterized, international populations.
Curtis Huttenhower: Even among well characterized clades, such as Bacteroides, we found genomes that were separate from characterized strains, so novel taxa without reference genomes.
Curtis Huttenhower: ASCA and ANCA antibody levels in blood corresponded significantly with microbial dysbiosis. We also found that each patient, and even some of the controls have a very dynamic microbiome, with different stable states, and periodic blooms.
Curtis Huttenhower: Our computational tools are available from the bioBakery website. (link: https://bitbucket.org/biobakery/biobakery/wiki/Home) bitbucket.org/biobakery/biob…
We also teach courses – and we are hiring computational and wet-lab postdocs, so please come and join our lab.
David Zeevi from Rockefeller University
The next speaker is David Zeevi, @DaveZeevi from Rockefeller University @RockefellerUniv with his talk entitled “Sub-genomic variation in the gut microbiome associates with host metabolic health”.
David Zeevi: What are we looking for in microbiome data? Observation? Association? Mechanism? We are moving from the first towards the last, but each step brings bigger challenges.
David Zeevi: Metabolic diseases are on the rise, in particular obesity. We collected blood glucose data and correlated that with microbiome and dietary data. Each person responds differently. See: (link: https://www.cell.com/cell/fulltext/S0092-8674(15)01481-6) cell.com/cell/fulltext/…
David Zeevi: Even small differences, in a few microbial genes, can have a significant phenotypic effect. Think about the presence/absence of a toxin gene. What are the variable regions in microbiome bacteria?
David Zeevi: About 20% of metagenomics read gets assigned to more than one reference microbial genome. Errors caused by differential coverage in reference genomes.
David Zeevi: We have a paper in press for an iterative coverage based algorithm for read-assignment correction. This creates more accurate assignments.
David Zeevi: Sub-genomic variability (SGVs). These regions are very abundant across different datasets (Elies: not sure how they are defined)
David Zeevi: SGVs are person-specific and are shared with habitat. SGVs also correlate with disease risk factors. E.g. Anaerostipes hadrus region: if present, people are leaner and more healthy. It encodes butyrate production and inositol degradation.
David Zeevi: Thus, variable gene clusters facilitate mechanistic insights. This would be not be discovered if you just look at presence of specific strains or species.
Kara Bortone from JLabs
After the break, we will have Kara Bortone @kara_bortone, head at jLABS @JLABS, Johnson & Johnson @JNJInnovation with “Catalyzing and supporting the translation of preventative research in the microbiome space”
Kara Bortone: Clinical research is still reactive: we wait until a person has a disease and then try to cure them. Can we study underlying mechanisms and better predict and prevent?
Kara Bortone: Future models of care: a holistic approach to eliminate diseases. Prevent, intercept, and cure disease. We are shifting our effort to a much earlier in the process, before the onset of observable symptoms.
Kara Bortone: The microbiome is a promising target for drug development. @JNJInnovation
has a recognized leading partnership role in the microbiome space. @JLABS
is an incubator space to grow and foster startup companies.
Kara Bortone: Jlabs are life-science incubators. There are now13 JLabs sites all across the globe, 480 portfolio companies in sectors ranging from health tech, medical devices, and pharmaceutical.
Kara Bortone: Small companies get big company benefits at JLabs. It is hard to buy equipment for early stage companies. So we have shared equipment and shared space, with support staff, education, and connections to VCs and other investors.
Kara Bortone: We organize events with investors and foundations to facilitate connections and QuickFire Challenges, with currently over 60 winning companies. And @AstarteMedical is one of them, yay!
Kara Bortone: Of the companies we hosted, 88% are still in business or acquired (that is more than the average restaurant in DC!). 12 companies have gone public, 12 were acquired.
That completes my tweeting from the Kisaco Microbiome Congress in Washington DC. I hope you all enjoyed this report!

















 Introduction
Introduction